Research Highlights
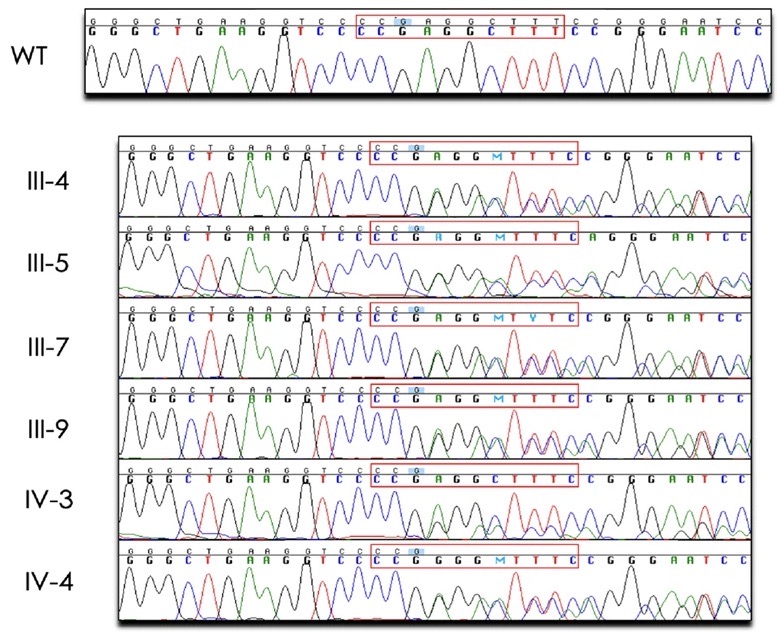
A dominant negative ADIPOQ mutation in a diabetic family with renal disease, hypoadiponectinemia, and hyperceramidemia
Adiponectin is an adipocyte-derived hormone with pleiotropic actions that promote insulin sensitivity, inhibit cell death, and decrease inflammation. Adiponectin forms an obligate trimer and circulates as trimers, hexamers, and high-molecular weight multimers that target various tissues and cell types, including liver, kidney, cardiac myocytes, and pancreatic β cells. Levels of adiponectin are decreased in obesity and may contribute to a chronic state of inflammation that leads to insulin resistance, type 2 diabetes, coronary artery disease, myocardial infarction, nonalcoholic steatohepatitis, and kidney disease. Recently, the antiapoptotic, insulin-sensitizing, glucose lowering, anti-inflammatory, and anti-steatotic effects of adiponectin have been linked to its role in sphingolipid metabolism and its receptor-mediated activation of ceramidase activity, which reduce levels of lipotoxic ceramides.
Previous studies have identified common variants in or near ADIPOQ, the gene that encodes adiponectin, that are associated with aberrant adiponectin levels, obesity, type 2 diabetes, and diabetic kidney disease. Rare ADIPOQ amino acid substitution mutations have also been reported in individuals with diabetes and hypoadiponectinemia. Here, we describe the first multigenerational family with a protein-truncating mutation in ADIPOQ (p.Gly93GlufsTer73), diabetes, and end-stage renal disease. Carriers of this mutation have dramatically reduced circulating adiponectin and increased long-chain ceramide levels. Functional studies show that this mutation acts via a dominant negative mechanism, with the wild-type and mutant proteins interacting, leading to decreased levels of cellular and secreted adiponectin. Our findings support adiponectin’s protective role and suggest that its genetic loss leads to ceramide accumulation which, in turn, contributes to diabetes and progression of renal disease.
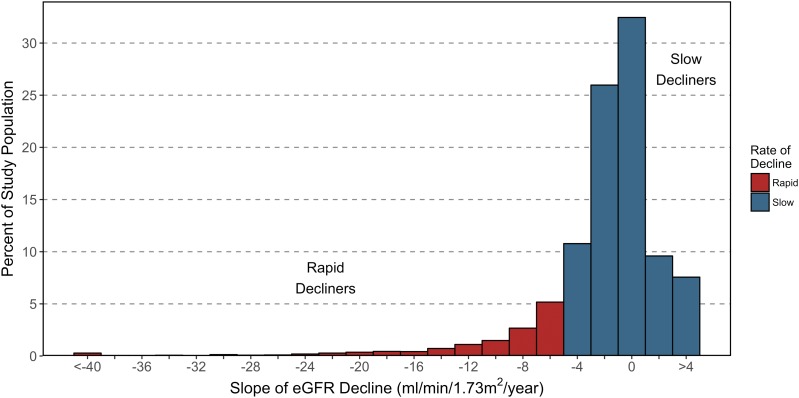
The Familiality of Rapid Renal Decline in Diabetes
Despite significant advances in the management of diabetes, including widespread implementation of renoprotective therapies when indicated, diabetic kidney disease (DKD) remains the leading cause of end-stage renal disease (ESRD) in the U.S. and is associated with excess morbidity and mortality.
Progressive glomerular filtration rate (GFR) decline precedes ESRD and has recently been established as the predominant clinical feature of DKD. For some patients with DKD, renal function declines gradually, whereas others experience a sudden decline, quickly reach ESRD (defined by a GFR <15 mL/min/1.73 m2), and require renal replacement therapy in the form of dialysis or the receipt of a functioning kidney through donor transplantation to survive. Despite interindividual variability, renal function decline progresses at a steady, or linear, rate over the course of DKD. In addition, those with more rapid renal decline have higher risk of all-cause mortality.
Wide individual variation in the rate of progression of renal function decline in diabetes has motivated recent studies to identify biomarkers that are associated with or predictive of rapid renal decline. Such biomarkers may have utility in patient surveillance and management. In addition to studies examining proteomic, metabolomic, and environomic profiles associated with renal decline in diabetes, various studies are also underway to identify genetic factors that contribute to its risk. Although these genetic studies are likely to be more powerful than previous studies that have examined a spectrum of DKD phenotypes, the familial risk of rapid renal decline has not been investigated. Understanding the familiality of rapid renal decline is paramount to recognizing the magnitude of the influence of shared factors, both environmental and genetic, on rapid renal decline.
To advance this area of research, we set out to investigate the familial nature of progressive renal function decline in diabetes using the Utah Population Database (UPDB), a unique research resource that links genealogical data to electronic health record data. Through this population-based retrospective cohort study, we show that there is strong evidence of familial clustering of rapid renal decline. Our findings support the hypothesis that shared familial factors (e.g., shared environmental and genetic factors) play a role in its susceptibility.
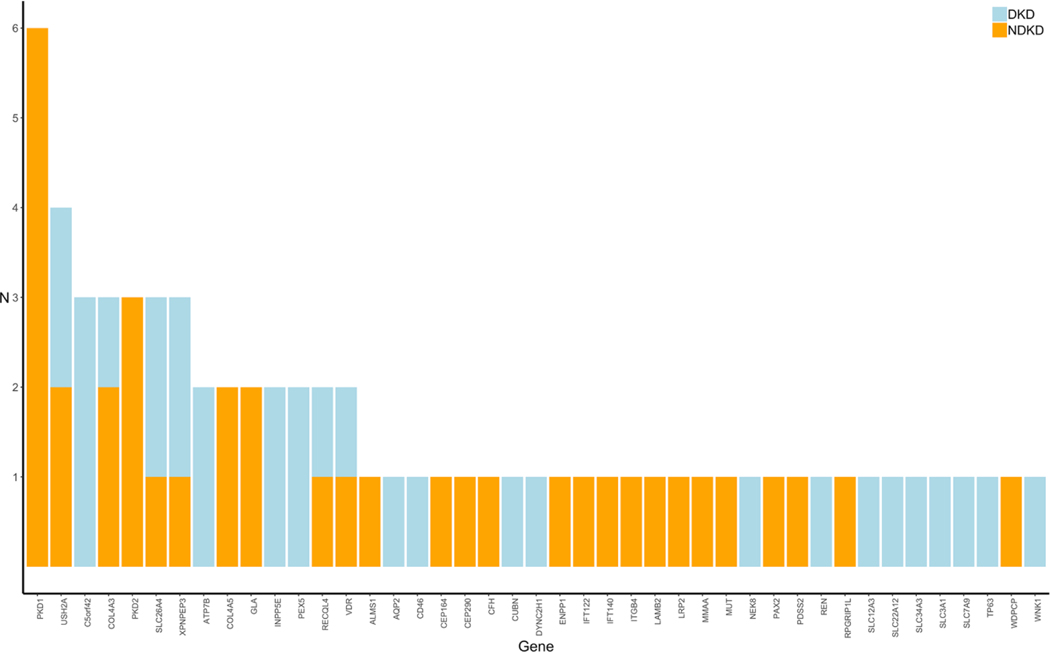
Targeted Next-Generation Sequencing Identifies Pathogenic Variants in Diabetic Kidney Disease
Diabetes is the most common cause of chronic kidney disease (CKD). Continued increase in the world-wide prevalence of diabetes has led to an increase in the global prevalence of diabetic kidney disease (DKD), also known as diabetic nephropathy. For patients with diabetes and CKD, the underlying cause of their kidney disease is often assumed to be a consequence of their diabetes. However, without histopathological evidence, it’s unclear whether such patients have true DKD, non-diabetic kidney disease (NDKD), or concomitant DKD and NDKD.
The only way to truly determine whether CKD in patients with diabetes is a consequence of diabetes is to perform kidney biopsies. Unfortunately, kidney biopsies are not clinically indicated in the diagnosis of DKD. Interestingly, recent investigations of kidney biopsies from patients with type 1 diabetes (T1D) or type 2 diabetes (T2D) and CKD have shown that as many as 30–83% of patients diagnosed with DKD actually had kidney disease attributed to non-diabetic causes. Familial focal segmental glomerulosclerosis (FSGS), followed by hypertensive nephropathy, acute tubular necrosis, and IgA nephropathy were the most common diagnoses observed in diabetic patients found to have NDKD. Importantly, accurately identifying a patient’s primary cause of CKD is a crucial component of its proper classification, prognosis, and management.
Recent studies conducted primarily in patients with NDKD have shown that next-generation sequencing (NGS) provides a promising avenue toward uncovering and establishing precise genetic diagnoses in various forms of kidney disease. The diagnostic yield from these studies range from approximately 10–40% and are highest among patients with congenital or cystic kidney disease. While these studies all show the utility of NGS in providing a molecular diagnosis for patients with heritable forms of kidney disease, whether this technology could also aid in the genetic diagnosis of patients with DKD is unclear. Importantly, improved understanding of the underlying disease process in DKD could have major implications in terms of patient care and monitoring as well as for research studies in this field. To date, no study has examined the distribution of rare variants in known kidney disease-related genes in unrelated patients with DKD. To address this, we performed targeted NGS using a custom gene panel comprised of 345 kidney disease-related genes to interrogate variations between identified NDKD and DKD to identify underlying causes of kidney complications in these patients.
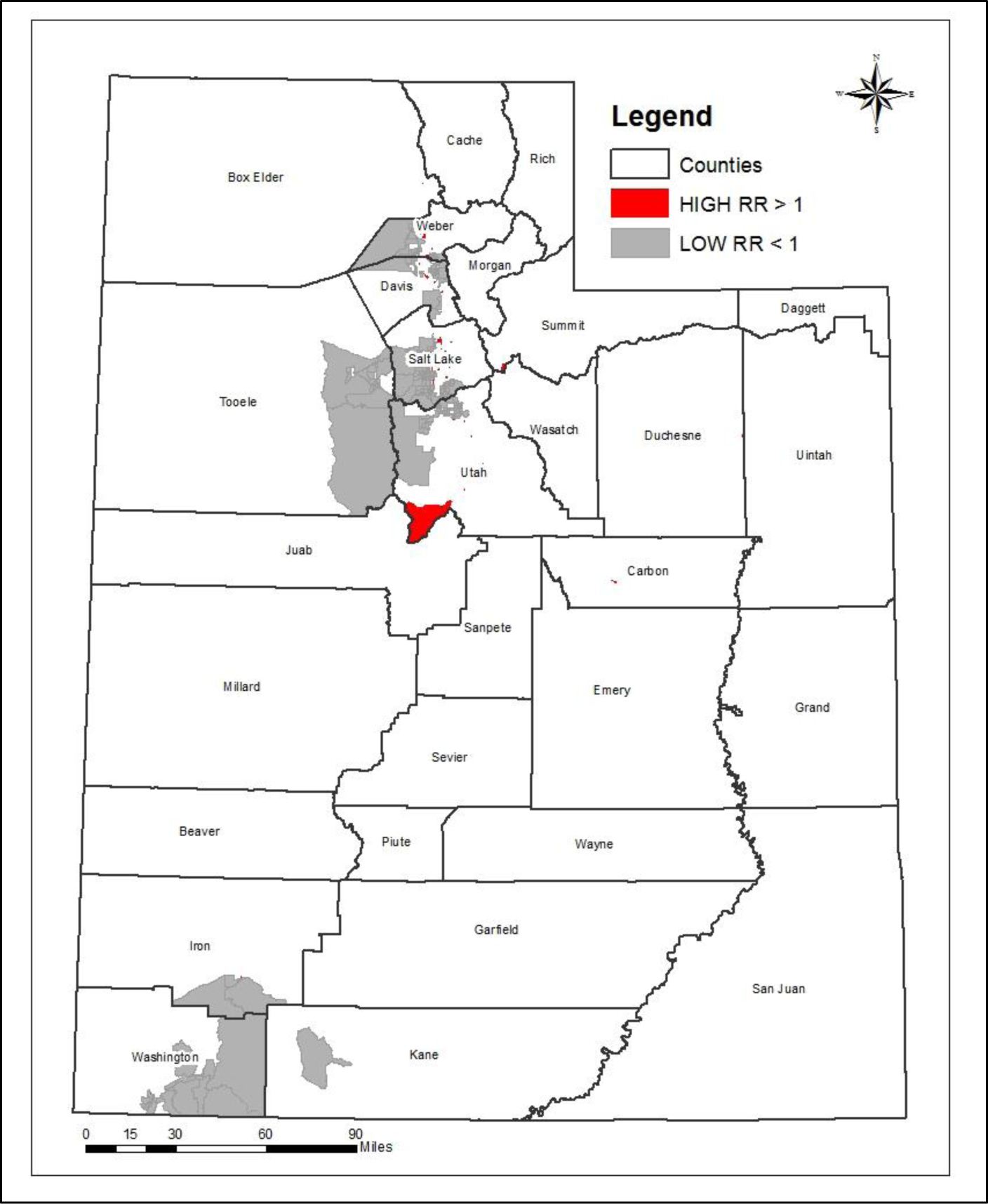
Type 1 Diabetes Incidence Among Youth in Utah: A Geographical Analysis
Type 1 Diabetes (T1D) is a unique and complex autoimmune disease that is heavily influenced by non-genetic factors such as socioeconomic status and environmental triggers. T1D is differentiated from Type 2 Diabetes (T2D) in that it is not caused primarily by dietary or lifestyle behaviors. In individuals with T1D, the body does not produce insulin and cannot, therefore, break down the sugars and starches consumed. A great deal of research has been done on the clinical aspects of T1D and the complications that result from it. Still, much less has aimed to identify the socio-demographic and environmental triggers associated with the spatial pattern of T1D incidence, as we do here. This research is unique in that it leveraged individual-level medical records from the Utah Population Database and analyzed the spatial patterns of T1D at a very fine geographic scale, the Census Block Group (CBG) level, in the state of Utah. One of the primary limitations of others papers has been the use of much larger scale geographic areas and the aggregation of data, which tends to hide spatial patterns. The primary contribution of this research was to identify hotspots or clusters at the CBG level, where the risk of developing T1D is significantly higher than the general population, and to clarify certain predictors of those hotspots.
T1D most often affects youth between the ages of 0–18. The highest rates of T1D occur between the ages of 5–7 and at or near puberty. T1D has been termed juvenile-onset diabetes, but it can occur at any age. The rate of diagnosed T1D has risen substantially over the past 30 years. The incidence trend worldwide is increasing at a rate of 3% annually. Finland has the highest incidence rate of T1D in the world, with 60 cases per 100,000 people each year, whereas China has a very low incidence rate of 1.01 per 100,000. The incidence rate of T1D among U.S. non-Hispanic white youth is 23.6 cases per 100,000 person-years. In Colorado, a state with a similar elevation as our study site Utah, the incidence rate reported in 2002–2004 was 23.9 per 100,000 person-years.
The cause of T1D is unknown but is believed to be a combination of genetic and environmental factors. Astonishingly, 85% of new cases of T1D have no family history of the disease.
T1D and T2D have some common characteristics such as high blood glucose levels that can have life-threatening complications, but they also differ in many ways. In T1D, the body’s immune system is responsible for attacking the beta cells of the pancreas and destroying them so that they no longer produce insulin to moderate blood sugar levels. In T2D, the body continues to produce insulin but does not use it properly (termed insulin resistance). Because of the body’s inability to produce insulin, T1D patients must receive insulin injections multiple times per day for the remainder of their life following clinical diagnosis.
A number of dietary and lifestyle factors have been considered to be triggers of T1D, including the early consumption of gluten and cow’s milk; vitamin D deficiency, as a result of higher latitude and reduced exposure to ultraviolet radiation; viral infections; and excessive hygiene. Research has shown that the incidence of T1D is positively related to distance north of the equator. Seasonal and temporal changes have also been found to influence the incidence of T1D, with more cases being diagnosed in the fall and winter months. Seasonal variations in the diagnosis of T1D have been linked to seasonal variations of certain viral infections, and have been documented as far back as 1926. These seasonal changes, along with the fact that very few cases have any family history of the disease, lead to the conclusion that certain unknown environmental factors are a crucial contributor to the disease process. This paper contributes to the body of research done on T1D by seeking to identify these unknown environmental factors that contribute to the incidence of T1D at the CBG level, with a focus on socio-demographic and geographic risk factors.
Socioeconomic status (SES) plays a vital role in regard to health behaviors and health outcomes. Many causes of mortality and morbidity are influenced by one’s income, education, and occupation. Research has shown that the association between SES and health outcomes typically shows a gradient pattern with improved health and decreased mortality with each increase in occupational grade (Adler and Ostrove, 1999). SES is often treated as a potential confounder between other variables and health outcomes (Braveman et al., 2005). Variations in SES across geographies often lead to differences in lifestyle, maternal age, diet, and exposure to infections, all of which may play a role in similar patterns of incidence of T1D among populations (Patterson and Waugh, 1992). Comparisons of the impact of SES on T1D are difficult, due to the fact that different definitions of SES are used as well as at different geographic scales. Little research has been done on the association between T1D incidence and SES at a fine spatial scale. To the best of our knowledge, only one study (Haynes et al. 2006) has assessed the relationship between T1D and SES for small area units (i.e., Australian census collection districts with approximately 250 dwelling per unit). The current study seeks to build on this knowledge gap by examining the relationship of T1D incidence and SES at the CBG level in Utah.
The primary purposes of this study are to calculate incidence rates of T1D among youth in Utah, examine the spatial pattern of cases, and measure the socio-demographic and geographic associations with the spatial patterns.